Research
1. Receptor Protein-Tyrosine Phosphatases (RPTPs): Dimerization, Substrate Interaction and Activity Modulation.
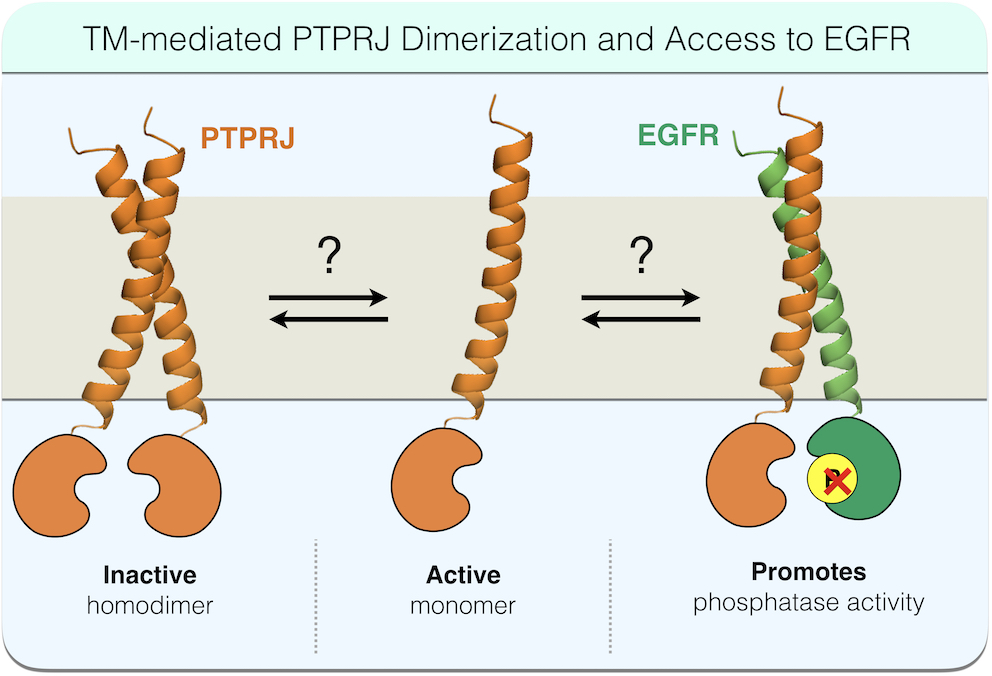
Receptor protein tyrosine phosphatases (RPTPs) play critical
signaling regulatory roles in development, health, and disease
progression. Despite their clear importance in signal
transduction, very little is known about the structure-function
relationships that underpin the regulation of their activity. The
reported ability of RPTP homodimerization to antagonize their
catalytic activity, however, presents potential opportunities to
develop therapeutic strategies to promote RPTP activity against
their oncogenic receptor tyrosine kinase (RTK) substrates.
For instance, our recent work, using PTPRJ/EGFR as a model RPTP/RTK pair, demonstrates that (homodimerization of RPTPs is regulated at least in part by transmembrane domain interactions, and disrupting these interactions can antagonize RPTP homodimerization, reduce substrate RTK phosphorylation, and ultimately inhibit RTK-driven cell phenotypes, including migration. While these results are exciting, much more work is needed to (1) determine the structural and cell dynamic basis for the observed effects, (2) develop realistic methods for therapeutically leveraging this new understanding, and (3) determine how different cellular contexts predict the outcome of interfering with PTPRJ dimerization through TM domains. To achieve these goals, we have been working in close collaboration with the Lazzara Lab (Chemical Engineering, University of Virginia) and the White Lab (Koch Institute for Integrative Cancer Research at MIT).
Ultimately, our studies stand to advance our basic biological understanding of RPTP biology and to lead to new methods to target signaling through oncogenic RTKs that may be less susceptible to common mechanisms of acquired resistance. This work is funded by NIH-NIGMS R01 award (GM139998).
2. Developing novel strategies to specifically target and deliver therapeutics to cancer cells and tumors.
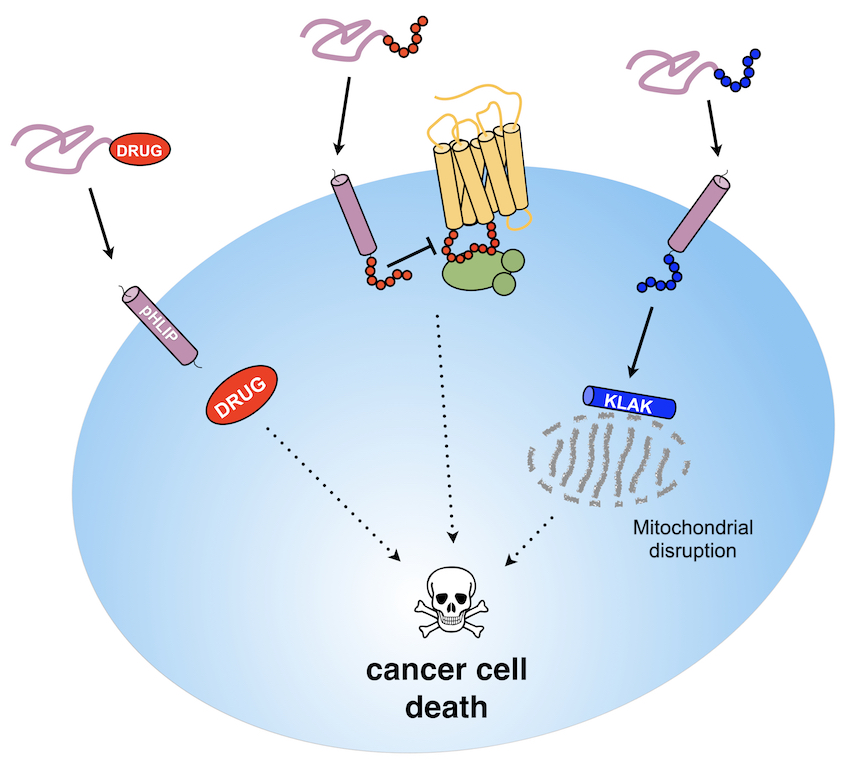 Preferential
targeting and delivery of chemotherapies to tumors have emerged as
a way to increase selectivity towards tumors, diminish the side
effects associated with uptake of drugs into healthy tissues, and
improve efficacy by targeting markers (i.e., proteins) present at
the surface of cancer cells. Biomarker over-expression provides a
narrow window of selective targeting but labeling of normal
tissues is also observed, which can lead to unacceptable toxicity
profiles. Second, therapy strategies based on the targeting of
specific proteins is significantly hampered by tumor
heterogeneity, which can promote tumor evolution, lead to loss of
cell surface proteins, and eventually result in drug resistance.
We have a uniquely designed cancer drug targeting and delivery
strategy, which has the potential to address both of those
critical shortfalls.
Preferential
targeting and delivery of chemotherapies to tumors have emerged as
a way to increase selectivity towards tumors, diminish the side
effects associated with uptake of drugs into healthy tissues, and
improve efficacy by targeting markers (i.e., proteins) present at
the surface of cancer cells. Biomarker over-expression provides a
narrow window of selective targeting but labeling of normal
tissues is also observed, which can lead to unacceptable toxicity
profiles. Second, therapy strategies based on the targeting of
specific proteins is significantly hampered by tumor
heterogeneity, which can promote tumor evolution, lead to loss of
cell surface proteins, and eventually result in drug resistance.
We have a uniquely designed cancer drug targeting and delivery
strategy, which has the potential to address both of those
critical shortfalls.
We are also highly involved, in collaboration with the Ladokhin Lab (University of Kansas Medical Center) in characterizing the interplay of lipid properties, pH and divalent cations in modulating the membrane interaction and tumor-targeting action of pH-sensitive transmembrane peptides such as pHLIP.Our group has been instrumental in showing that pHLIP can be developed as as a versatile and efficacious targeting and delivery platform to improve cancer therapy modalities. See Publications. For instance, we have shown that pHLIP provides localized delivery of monomethyl auristatin derivatives not only to cultured cancer cells but also to tumors in mouse models. We recently reported the first-in-class pHLIP-based immune-engaging agents that directed the recruitment of antibodies to cancer cell surfaces. This work is funded by two NIH-NCI R21 awards (CA259800 and CA267087).
3. How do Cells Sense and Decode Flow
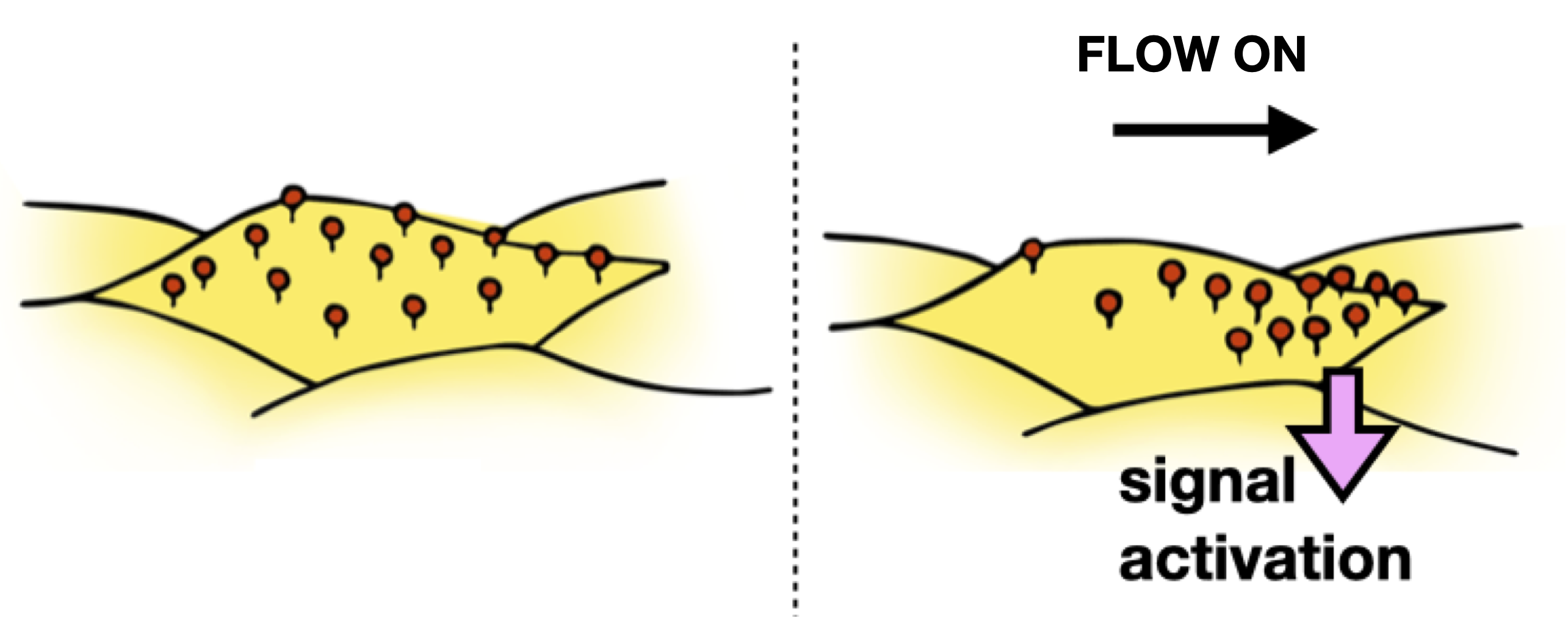 These
studies in collaboration with the Honerkamp-Smith
Lab (Physics, Lehigh University) are aimed at quantitatively
define and predict how endothelial cells sense, interpret, and
respond to shear flow.
These
studies in collaboration with the Honerkamp-Smith
Lab (Physics, Lehigh University) are aimed at quantitatively
define and predict how endothelial cells sense, interpret, and
respond to shear flow.
Our approach is to conduct parallel experiments in model membranes and living cells, allowing us to directly relate physiological function to molecular biophysics, and intracellular cell signaling. While the model protein studied is specific to endothelial cells, the principles of fluid mechanics that we will uncover are universal. We, therefore, anticipate that our models will apply to multiple cell lines and flow conditions, and will lay the groundwork for future research directions. This work is funded by by NIH-NIGMS R01 award (GM143320) and a New Initiative Research Award from the Charles E. Kaufman Foundation.Our central hypothesis is that physiologically significant protein and lipid concentration gradients arise from physical interactions between fluid flow and complex membranes. This hypothesis is based on the premise that extracellular lipid-anchored proteoglycans like glypican-1 can be transported along the plasma membrane by external flow, with the aqueous part of the protein acting as a molecular sail.