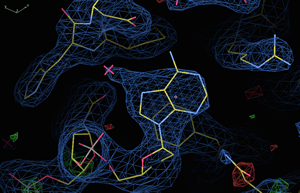
 Researchers in the McLaughlin Lab use the tools of structural biology to determine the molecular structure of proteins and protein complexes. Structural biology coupled with molecular biology, biochemical, and biophysical techniques allow us to investigate structure-function relationship in various systems.
Researchers in the McLaughlin Lab use the tools of structural biology to determine the molecular structure of proteins and protein complexes. Structural biology coupled with molecular biology, biochemical, and biophysical techniques allow us to investigate structure-function relationship in various systems.
The McLaughlin Lab has multiple research projects in progress.
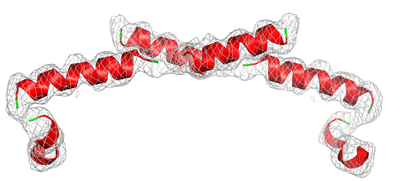
A major research interest of our lab is the the study and characterization of protein targets from the commensal human gut bacteria Bacteroides ovatus. Gut microbiota populations in healthy individuals show marked differences compared with populations in individuals with autoimmune diseases. An overabundance of the normally commensal gut microbe Bacteriodes ovatus has been observed in the autoimmune disorders ulcerative colitis, Crohn’s disease, type I diabetes and celiac disease. We will use x-ray crystallography coupled with several biochemical and biophysical techniques to study the microbe Bacteriodes ovatus. We are performing functional and structural investigations of protein targets from B. ovatus hypothesized to play a part in provoking a human autoimmune response or on strain survival.
 |
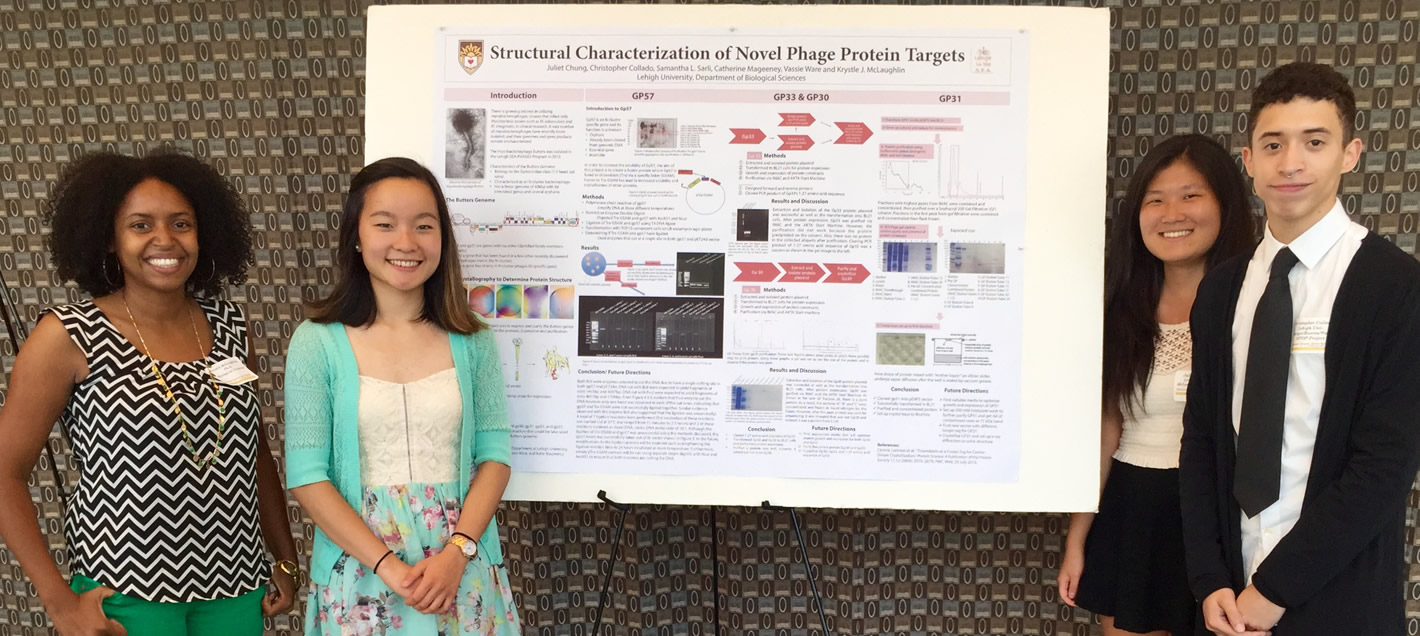
The other focus of the lab involves a collaboration with Dr. Vassie Ware and the Lehigh SEA-PHAGES program. We are structurally characterizing novel protein targets from bacteriophages identified by Lehigh undergraduates students in the Lehigh in the S.E.A course.
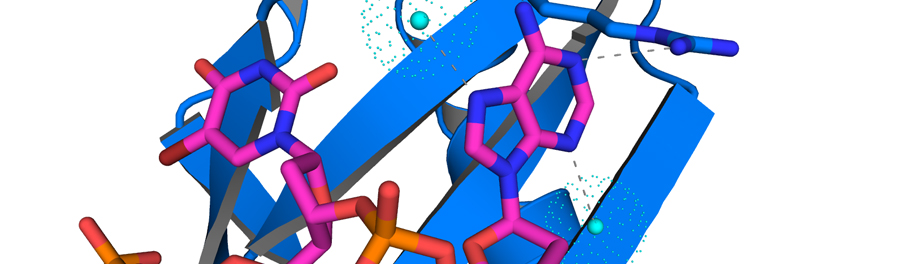
Our primary research tool is x-ray crystallography, and we also employ biochemical and biophysical methods such as recombinant protein purification, fluorescence anisotropy and enzyme-linked assays to study various protein targets.
 |
- Lysozyme, Models and the PDB: Healping Undergraduates Explore Structure (Protein Data Bank, Spring 2016)
Agrawal A., McLaughlin K.J., Jenkins J.L., Kielkopf C.L. “A Structure-Guided U2AF65 Variant Improves Recognition and Splicing of a Defective pre-mRNA.” Proc Natl Acad Sci U S A. 2014, 111(49):17420-5
McLaughlin K.J., Nash R.P., Redinbo M.R. “Unique Helicase Determinants in the essential conjugative factor TraI from Salmonella typhimurium plasmid pCU1.” J. Bacteriol. 2014, 196(17):3082-90 doi:10.1128/JB.01496-14 Pubmed
McLaughlin K.J., Jenkins J.L., Kielkopf C.L. “Large Favorable Enthalpy Changes Drive Specific RNA Recognition by RNA Recognition Motif Proteins.” Biochemistry 2011, 50(9):1429-31 Text/PDF
McLaughlin K.J., Strain-Damerell, C.M., Xie, K., Brekasis, D., Soares, A.S., Paget, M.S. and Kielkopf, C.L. Structural Basis for NADH/NAD+ Redox Sensing by a Rex-Family Repressor. Molecular Cell. 2010 May 28;38(4):563-75. Text/PDF
Buboltz JT, Bwalya C, Williams K, Schutzer M. “High-resolution mapping of phase behavior in a ternary lipid mixture: do lipid-raft phase boundaries depend on the sample preparation procedure?” Langmuir 2007, 23:11968-71. Text/PDF
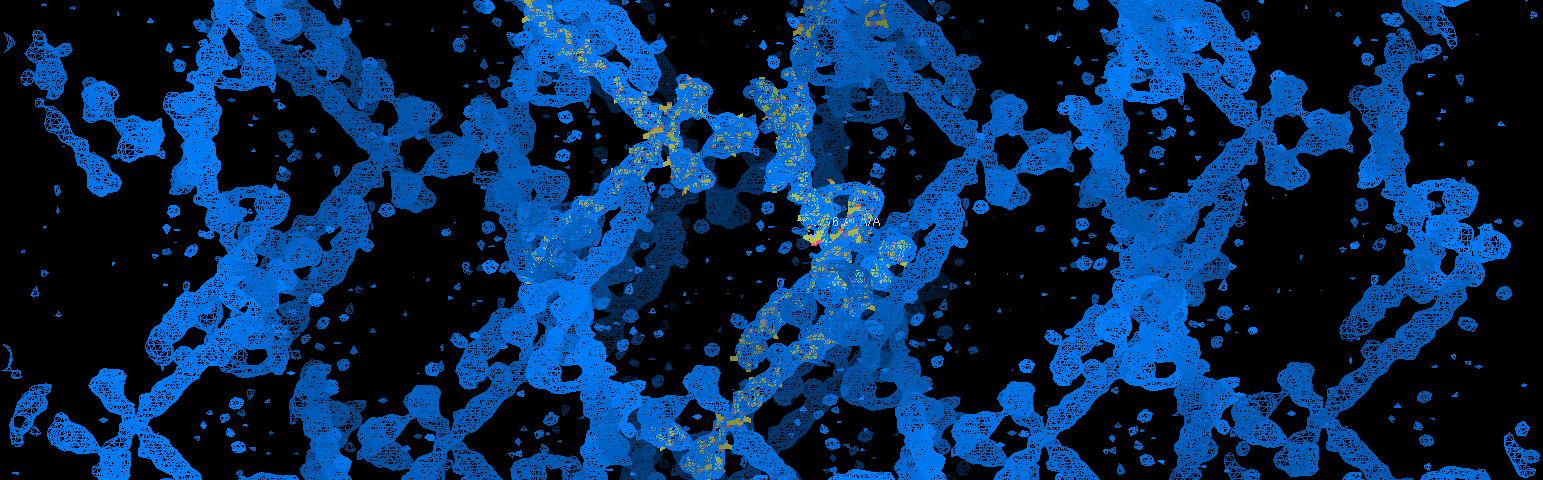

- "Structures of Phages"- Acumen, Lehigh University College of Arts and Sciences magazine. April 2015


